Fastidious plant diseases and plant-microbe interactions

Potato Zebra chip

Citrus greening
Fastidious (unculturable) plant pathogens are devastating to several food and commodity crops. For instance, citrus greening or Huanglongbing disease, caused by a fastidious bacterium, Candidatus Liberibacter asiaticus, is inflicting approximately $3 billion in annual losses. Similarly, potato zebra chip disease, caused by Candidatus Liberibacter solanacearum, causes annual crop losses of US $25 million in Texas alone. We are utilizing the latest genomic, genetic and biotechnology tools to discover and utilize novel disease resistance genes and antimicrobials (small molecules, peptides, etc.) to confer tolerance to the devastating pathogens. Furthermore, to overcome the challenges in studying fastidious pathogens, we are developing new technologies and bioassays that enable culturing and propagation of these pathogens. These tools are being used to conduct high throughput screening of antimicrobial genes and therapeutics.
FUNDING: USDA-NIFA-CDRE; Foundation for Food and Agricultural Research, Southern Gardens Citrus
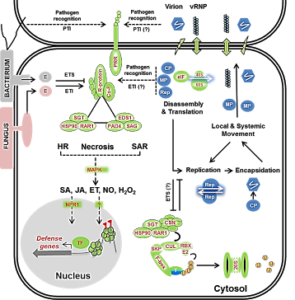
Monocot stress biology and grass-virus interactions
Diseases and abiotic stresses of grasses result in annual yield losses of US $300 million or more. However, very little is known about the gene regulatory networks that function in grass stress responses, particularly related to grass viral diseases. To enable fundamental studies of cereal and bioenergy grass defense pathways, we are pursuing genetic and genomic studies using model grasses, Brachypodium distachyon (a C3 grass) and Setaria viridis (a C4 grass). Using these models and employing latest omics technologies, we aim to speed discovery of grass immune signaling components. We identified the transcriptome-, spliceome- and metabolome-level changes occurring during diverse grass:viral infections. From these studies, we identified approximately thirty receptor-like kinases (RLKs) in several sub-families including multiple pattern recognition receptors (PRRs), defense hormone-related genes, reactive-oxygen species (ROS) homeostasis genes, transcription factors (WRKY, MYB and AP2/ERF family), protein synthesis- and degradation-related proteins that are mis-expressed. Each of these proteins could influence plant immune responses. Currently, we are characterizing how defense hormone salicylic-acid homeostasis is perturbed during virus infection and dissecting the role of Brachypodium PHENYLALANINE AMMONIA-LYASE (PAL) promotes antiviral defenses genes using genetic, molecular and biochemical approaches. The research findings are further being used to confer tolerance of agronomic grasses (sugarcane and energycane) to biotic and abiotic stresses using biotechnology and breeding tools.
FUNDING: USDA-NIFA-AFRI; DOE-JGI
Mechanisms of plant central stress regulators

BT2 is a plant central stress regulator. Mandadi et al., 2009, Plant Physiol.
Central stress regulators are an emerging signaling concept, where in few core genes respond to, and integrate multiple stress signals to promote tolerance against diverse stresses. In Arabidopsis, we characterized a central stress regulator, BT2, which mediates responses to diverse biotic and abiotic stresses, nutrient, environment and hormone signals [Mandadi et al., Plant Physiology 150 (2009), Plant Cell 19 (2007)]. We are currently pursuing functional characterization of BT2 signaling network in Arabidopsis.The BT2 signaling pathway is composed of other proteins including CULLIN3, GTE9/BET9 and GTE11/BET10 (Misra et al., 2018), as well as, BT2 homologs, BT1, BT3, BT4 and BT5. BT family proteins, including BT2, are hypothesized to assemble into distinct ubiquitin ligases that target specific proteins for ubiquitin-mediated proteolysis. GTE9 and GTE11 are also putative acetyl-histone binding bromodomain proteins, and they physically interact with BT2 in yeast 2-hybrid and in vitro co-IP interaction assays. BT2 also suppresses DNA methylation and promotes transcription mediated by viral enhancers. It is possible that the BT2 ubiquitin ligase targets specific proteins in transcription processes such as transcriptional activators, repressors and/or chromatin remodeling factors for degradation by polyubuiquitilation, and may influence DNA methylation status.
We are currently determining the role of BT2 complex in transcriptional regulation and interactions among the BT-family ubiquitin ligase components using genetic, biochemical and cell-biological techniques. This will further our fundamental understanding of the intricate cross-talk occurring between multiple abiotic and biotic stress signaling pathways, as well as discovery of novel components useful for crop improvement. The research findings from model plants are further translated to agronomic crops such as tomato, potato and citrus using biotechnology and breeding tools.
FUNDING: USDA-NIFA-HATCH, Texas A&M AgriLife Research